Genetically Engineered Crops
The knowledge of naturally occurring intra- and interspecies gene transfer events, the advances in recombinant DNA technology, and plant tissue culture helped scientists transcend traditional breeding limits and introduce desired genes from any organism into crops. The crops containing one or more genes from other species (trans/foreign gene) are known as genetically engineered/modified (GE/GM) crops.[1] For example, many insect-resistant GE crops contain the delta-endotoxin gene from a bacterium, Bacillus thurigiensis (Bt).
The purpose of making transgenic crops could be to increase the crop yield, prevent damage from pests and pathogens, and increase the nutritional quality. In 1994, the Food and Drug Administration (FDA) approved the first GE crop, Flavr Savr tomato, for human consumption. This tomato variety has a longer shelf life due to delayed fruit ripening. Since then, the FDA has approved more than fifty GE crops.[2]
This chapter summarizes the advances made in genetics and other related technology that led to the development of genetically engineered crops and provides examples of a few success stories.
7.1 DNA Is Genetic Material
In the early 1950s, it was proven that DNA is genetic material. In 1954, Francis Crick and James Watson proposed the double-helix model of DNA based on evidence from Morris Wilkins’s and Rosalind Franklin’s experiments. They suggested that DNA is a double-stranded, helical structure made of four nucleotide bases: adenine (A), thiamine (T), cytosine (C), and guanine (G). The two strands of DNA are antiparallel and are connected by covalent bonds: adenine from one strand binds to thiamine in another strand, whereas cytosine of one strand binds to guanine of the second strand (see figure 7.1). Thus the nucleotide sequence of one strand is complementary to another strand. If the sequence of one strand is known, it is easy to deduce the second strand sequence. Watson and Crick explained that one strand could serve as a template for making the other strand due to their complementary nature. Thus DNA contains information for self-replication—a necessary qualifier for being genetic material. In 1962, Crick, Watson, and Wilkins received the Nobel Prize for the elucidation of DNA structure.
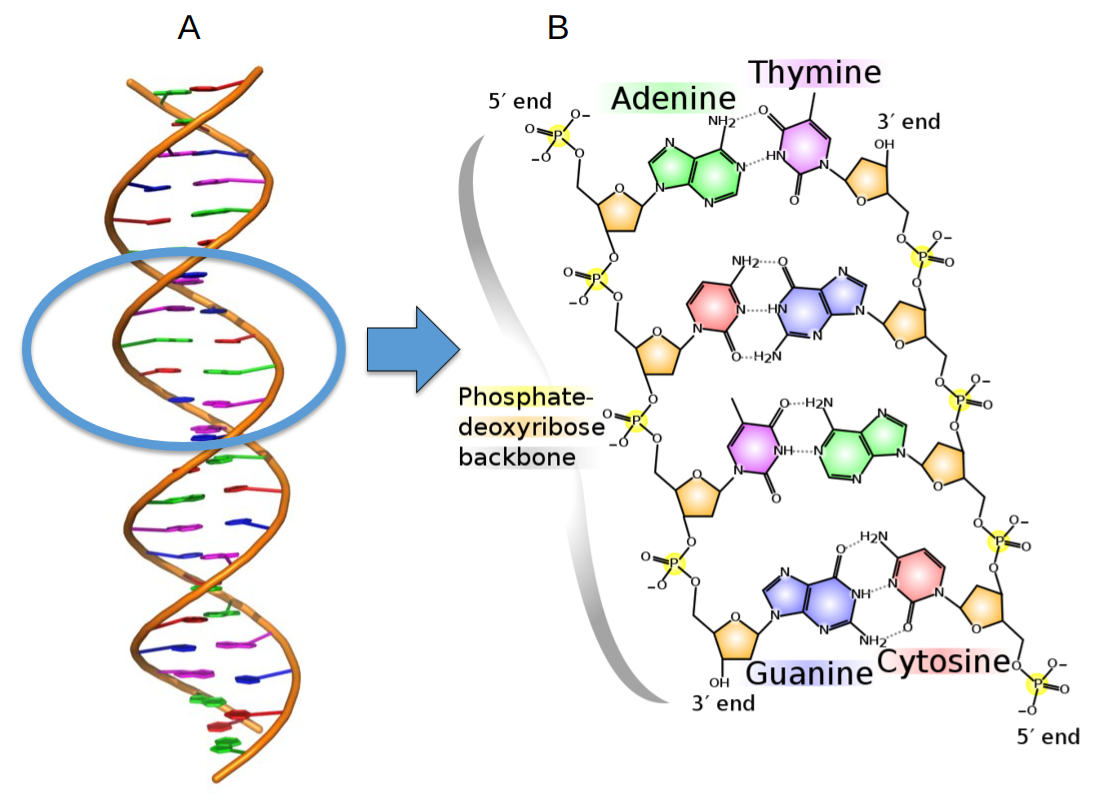
In 1958, Arthur Kornberg extracted a DNA polymerase enzyme from E. coli cells that was capable of copying DNA. He demonstrated the replication of a small DNA fragment in vitro using DNA polymerase and proved that either strand of DNA could serve as a template during DNA replication. DNA replication is a semiconservative process: four strands of DNA are made from two parental strands, and eventually, two double helixes form; each helix contains one old and one new strand. Other scientists’ research showed that DNA replication is not completely foolproof, and a proofreading and repair process ensures the removal of mismatched bases and repairs DNA. Thus DNA sequences are preserved for millions of years and ensure the continuity of a biological species. However, a permanent change in the DNA sequence (mutation) can occur due to the extremely low level of proofreading errors during the DNA replication. Most single nucleotide mutations are harmless and don’t affect a phenotype. However, after many generations, the accumulation of mutations could result in phenotypic changes. Thus DNA also acts as raw material for evolution.
The complete set of genetic material present in an organism is called its genome.[3] The genome of prokaryotes is made of a single circular chromosome, whereas the genome of eukaryotes consists of more than one chromosome. Individuals of the same species have the same genome size (same number of chromosomes), and chromosomes between different species vary. The structural and functional unit of the genome is a gene. The order of the four nucleotide bases determines the composition of individual genes. Several thousand genes are present within the organisms. Typically, the information from the DNA (gene) is transcribed into RNA, and a subset of RNA (messenger [m] RNA) is then translated into proteins. The proteins carry out most of the structural and functional activities within the cells. This unidirectional flow of genetic information from DNA to protein is called central dogma and serves as the fundamental principle for the basis of life. In addition to protein-coding mRNAs, organisms’ genomes contain genes that encode various other types of structural and regulatory RNA molecules (e.g., rRNA, tRNA, and microRNA).
7.2 Recombinant DNA Technology
After the 1970s, a new branch of biology, molecular biology, came into existence. Its focus is to understand the structure and functions of genes and gene products. It was expected that the sum of the knowledge about all genes could help in understanding the whole organism. The basic principles of molecular biology were discovered by experiments conducted on unicellular bacteria and fungi, and later studies on higher eukaryotes, including plants, began.
First, scientists succeeded in extracting DNA from various organisms. The next challenge was to identify the thousands of genes and then determine their function within the cell. The discovery of restriction enzymes and ligase enzymes made the identification and cloning of the individual genes possible. The restriction enzymes recognize a specific sequence of four/six/eight nucleotides within DNA and make a cut in the strand. The sites of restriction enzymes are randomly scattered within the genome, and hence they cut double-stranded DNA into small pieces. So far, more than 3,000 restriction enzymes have been discovered, of which 600 are available in the market. These enzymes were named after the bacteria where they were first identified. For example, the EcoR1 enzyme is found in the Escherichia coli strain RY13, and HindIII was discovered in Hemophilus influenzae.
Bacteria make restriction enzymes to protect themselves against virus infection. When viral DNA enters the bacterial cells, these restriction enzymes slice the virus DNA and block the virus’s growth. However, bacterial DNA remains protected from their own restriction enzymes by methylation of the corresponding restriction sites. In 1978, Werner Arbor, Hamilton Smith, and Daniel Nathans were jointly awarded the Nobel Prize for the discovery of restriction enzymes. In 1967, a special enzyme, ligase, capable of joining two DNA fragments, was discovered in Gellert, Lehmann, Richardson, and Hurwitz’s laboratories. For scientists, the restriction enzymes served as molecular scissors, and ligase served as a molecular glue for cutting and joining DNA fragments.
The next question was how to amplify small DNA strands for their detailed analysis. Here, the knowledge about a bacteriophage lambda that infects E. coli bacteria explicitly came in handy. Esther Lederberg discovered the bacteriophage lambda in 1951. Esther Lederberg observed that after infecting E. coli, lambda could undergo either an active lytic or a latent lysogenic cycle. When the bacterial cell grows in the rich medium (rapidly dividing), then the lambda enters the lytic cycle, makes 1,000 of its copies by using resources and machinery of bacterial cells, and then destroys the host. These 1,000 lambda offspring then infect 1,000 new cells and produce 1,000,000 new progenies in the next cycle. In this way, the virus grows at the speed of a rocket. However, if bacterial cells are deprived of nutrients and are in the stationary phase (not dividing), lambda enters the lysogenic cycle resulting in the insertion of the lambda DNA into the bacterial genome without causing any trouble. However, if the situation worsens for bacteria (or it experiences heat stress), lambda enters the lytic cycle to makes use of whatever resources are available for making its copies before host cells die. For lambda, going to a lytic or lysogenic state is controlled by the repressor protein encoded in lambda DNA. When lambda DNA enters the bacterial cell, first, the repressor protein is made, which blocks the rest of lambda’s genes. The rapidly dividing bacterial cells contain high levels of proteases, which can break down the repressor and allows the lambda to enter into the lytic cycle. Under nutritional deprivation, bacterial cells come to a stationary stage where the protease levels remain low, and the lambda goes into the dormant lysogenic cycle. When the host cells face stress (e.g., high temperatures), protease protein increases to mitigate the situation and recycle the nutrients. Thus in such a situation, the repressor protein is destroyed, and lambda enters the lytic cycle. Based on this knowledge of the lambda life cycle, scientists created versions of lambda in which researchers could easily turn the lytic and lysogenic cycle on or off. Furthermore, scientists found that only 75 percent of the lambda genome is indispensable; 25 percent of the lambda genome can be replaced with any DNA fragment. Thus if a gene is inserted within the lambda genome, millions of clones can be made quickly in E. coli. Then this DNA could be isolated, stored, and used for further experimentation. The lambda served as the first vector for cloning genes.
In the 1950s, Esther and Joshua Lederberg also observed that bacteria have many (10–1,000) small, circular DNA structures called fertility (F) plasmids in addition to a large genome. Before cell division, plasmids undergo autonomous replication and are inherited by the daughter cells. Soon, from a pathogenic bacteria, Shigella, another type of plasmid was discovered that contained antibiotic resistance genes (known as resistance [R] plasmids). It was easy to isolate plasmid DNA from bacterial cells, and also, bacterial cells have a natural capability of uptaking these small plasmids. The introduction of these plasmids into bacterial cells is called transformation. Scientists optimized protocols for increasing the efficiency of transformation and created chimeric plasmids by joining parts of F and R plasmids that contained one or two antibiotic-resistance genes, sites of restriction enzymes, and the autonomous origin of replication for E. coli. Thus these plasmids served as excellent vectors for cloning small DNA fragments. If a mixture of plasmid DNA and bacterial cells are plated in a medium containing a specific antibiotic, then the normal bacterial cells will die, and only transformed cells containing the desired plasmid (clone) will grow.
After the 1970s, the cloning of small DNA fragments (less than 20 kilobase pair [Kb]) in plasmid vectors and lambda-based phasmids became routine. Soon methods for DNA sequencing were invented, which made it possible to read the coded information (the order of four nucleotide bases, A, T, G, C) present within the genes. Computer analysis was used for analyzing sequencing data, and the field of bioinformatics was born. By comparing the genes present within living beings, it was found that the genetic code and genetic mechanisms are conserved among bacteria, plants, and animals. Computer algorithms are also used for assessing the similarity between the homologous genes from different organisms. The homologous genes between closely related species are more similar than distantly related species. In the late 1990s, due to advances in sequencing technology, it became possible to sequence whole genomes of bacteria, fungi, plants, and animals. Comparing the entire genome of different organisms allowed quantifying the similarities among the various organisms and deciphering the evolutionary distance between the species (see figure 7.2). In the light of modern genetics, molecular biology, and phylogenomics, we can now understand the process of biological evolution in reverse order.[4]
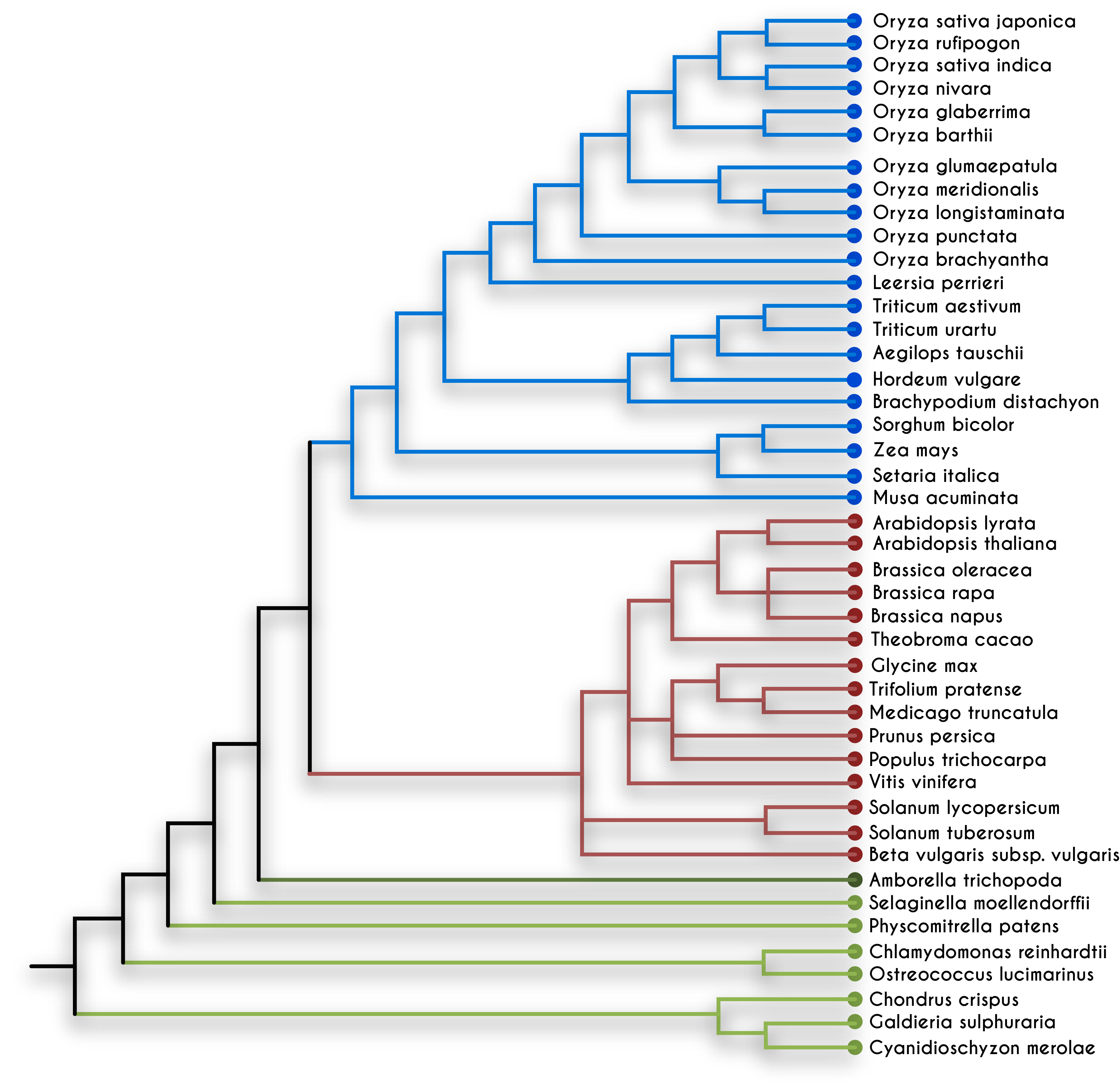
7.3 Gene Transfer in Plants
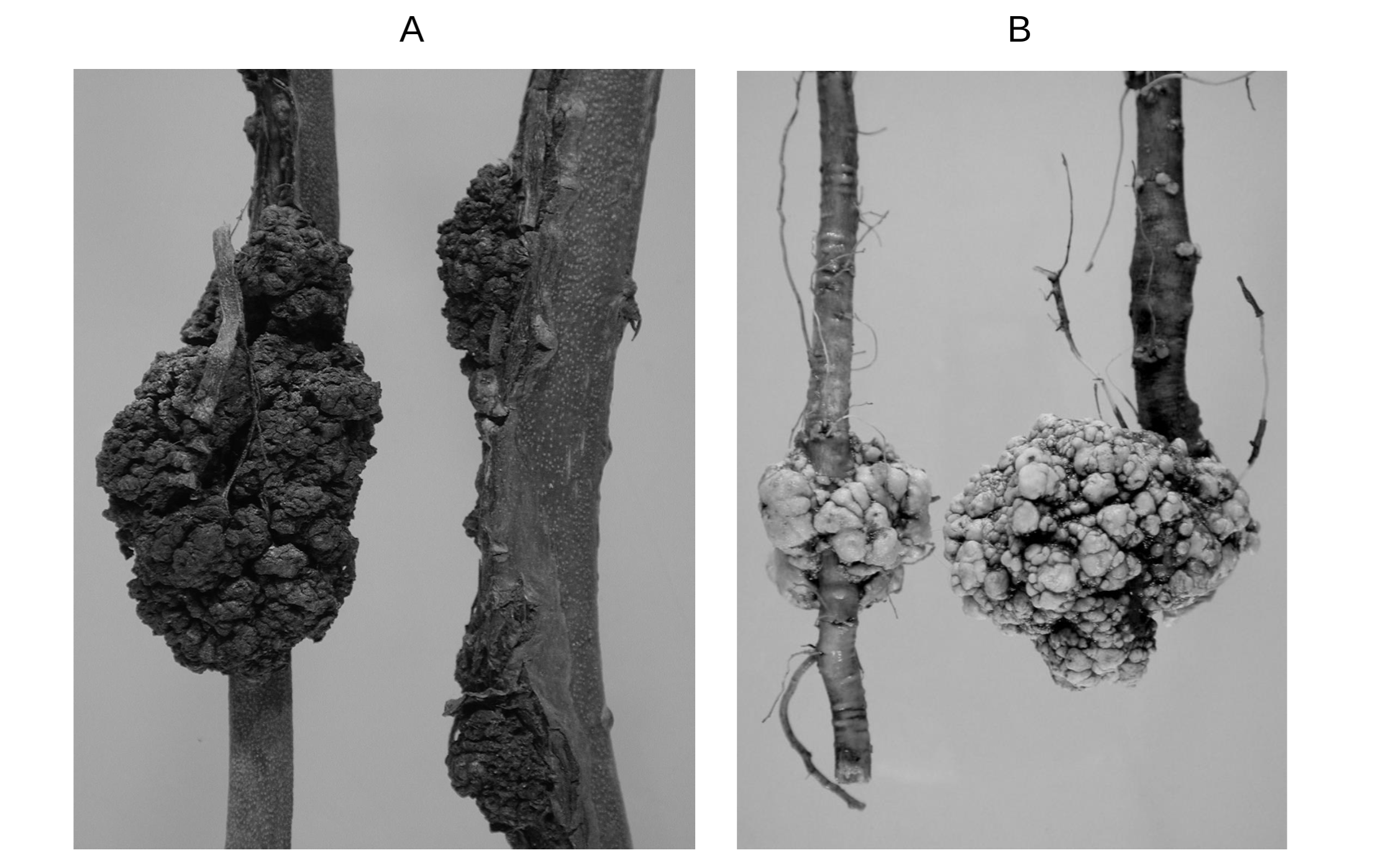
We often see swollen, round, tumor-like knots on the roots, twigs, and branches of many plants around us (see figure 7.3). These knots, called crown galls, are caused by a soil bacterium, Agrobacterium tumefaciens. These galls do not cause the death of the host plant but stunt their growth. The virulence of Agrobacterium strains is determined by a Ti plasmid present within the bacterium. Marc Van Montagu and his colleagues studied eleven pathogenic and eight nonpathogenic strains of Agrobacterium and found that all pathogenic strains had Ti plasmids and all nonpathogenic strains lacked Ti plasmids. They also noticed that the introduction of Ti plasmids into nonpathogenic strains transforms those into pathogenic strains. Thus it was concluded that Agrobacterium needs Ti plasmids for infecting the plants. The sequencing of the Ti plasmid revealed that it contains virulence (vir) genes and genes for the biosynthesis of auxin, cytokinins (plant growth hormones), and opines. The vir genes help bacterium infect host plants. The plant hormones promote the rapid growth of the cells at the site of infection (formation of crown galls), and opines serve as sources of energy, carbon, and nitrogen for Agrobacterium. Research in the 1970s and 1980s showed that Agrobacterium’s persistent presence within galls is not required for tumors to grow. In fact, after infecting a plant cell, Agrobacteria leave the Ti plasmid in the host cell, and then a large fragment of Ti plasmid, called T-DNA, gets inserted into the plant genome. The T-DNA in the plant genome continues to provide instructions for promoting the growth of the crown gall tumor and the biosynthesis of opines.
If a gene is inserted within the T-DNA of the Ti plasmid using recombinant DNA technology, it can be easily transferred within a plant genome using Agrobacterium. Scientists modified Ti plasmid by replacing genes responsible for tumor growth with restriction enzyme sites, antibiotic resistance genes, and the E. coli plasmid replicons. Thus chimeric Ti plasmids allowed cloning of foreign genes in them and were able to replicate within both E. coli and Agrobacterium. Scientists had the flexibility of efficiently cloning genes in E. coli and then use Agrobacterium for transferring the desired gene into the plant genome (plant transformation). Since Agrobacterium strains have a limited host range, they cannot transform all species of plants. The gene gun was used for transforming plant species, for which agroinfection was not an option. In this alternative method, very fine particles of gold or tungsten coated with desired DNA are bombarded on plant tissue by the gene gun. The advances in plant tissue culture made it possible to regenerate whole plants from small plant parts. Thus the combination of recombinant DNA technology and plant tissue culture allowed the bioengineering / genetic engineering of plants for important quality traits (that were not available in a given plant species’ diversity pool). It became possible to introduce useful genes from bacteria, animals, and other plant species into the crop species.
Both methods of plant transformation use small plant parts to start with, and after agroinfection or gene bombardment, the selection of transgenic cells is carried out in the synthetic tissue culture media (see figure 7.4). One or more antibiotic selection makers accompany the transgene; thus only transgenic cells grow in this media while normal cells die. After several rounds of selection and multiplication of transgenic cells in the tissue culture, tests for the desired transgene are carried out. Once the insertion of the transgene is confirmed, the expression analysis and other assays are conducted, and whole plants are regenerated.
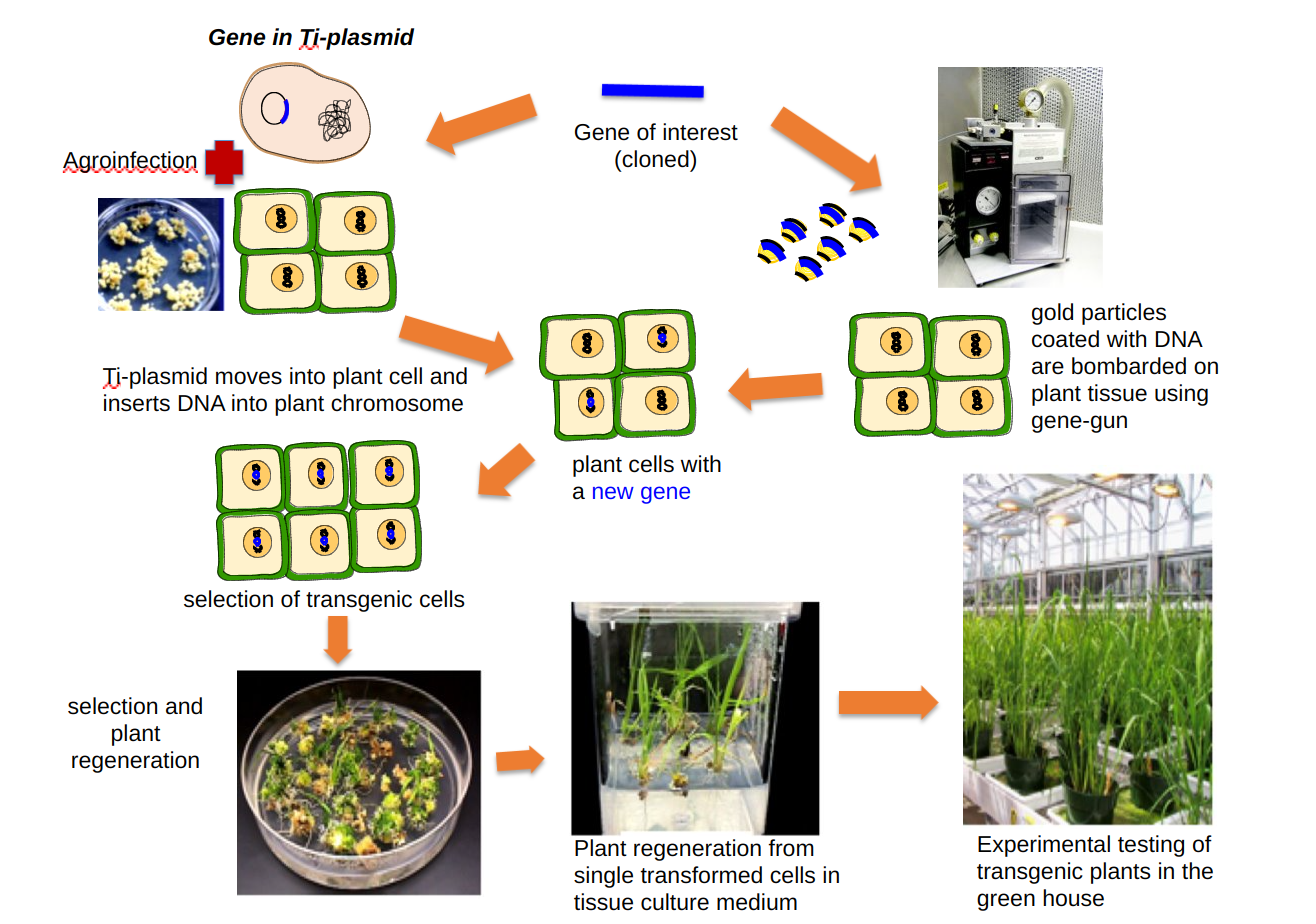
In general, both plant transformation methods have a low efficiency, and then among the transformed plant cells, only a few show the desired level of transgene expression. Once the transgenic plants are fully developed, the second round of testing begins, and one to five of the best-performing plants are selected.
7.4 GE Crops
The GE crops made so far can be divided into three classes: (i) pest or pathogen resistant, (ii) herbicide tolerant, and (iii) biofortified with improved nutritional value (see table 7.1). At present, ~15 GE crops are being grown in 12 percent (179.7 million hectares) of the total agricultural land worldwide. In the US alone, ten GE crops are being grown on 70 million hectares. Maize, soybean, and cotton are among the most grown GE crops. Other GE crops include apple, mustard, sugar beet, papaya, potato, pumpkin, eggplant, alfalfa, poplar, rose, golden rice, golden cassava, and so on.[5]
7.4.1 The Story of Virus-Resistant Rainbow Papaya
The delicious papaya fruit is a rich source of vitamins A and C, calcium, and potassium. Papaya (Carica papaya) is the native plant of Central and South America. When Christopher Columbus tasted it for the first time, he was overwhelmed and named it the “fruit of the angels.” After the fifteenth century, papaya spread throughout the world under the umbrella of European colonialism. Today it is grown in India, Brazil, the Philippines, Indonesia, Malaysia, Thailand, Hawaii, and the Caribbean islands. Usually, within six months after sowing the seed, the papaya grows as high as the average tree and starts bearing fruit within a year. Within three years, the papaya tree matures and produces a full yield. The cost of setting up papaya plantations is less than that of other plantations, and the farmers make decent profits from it.
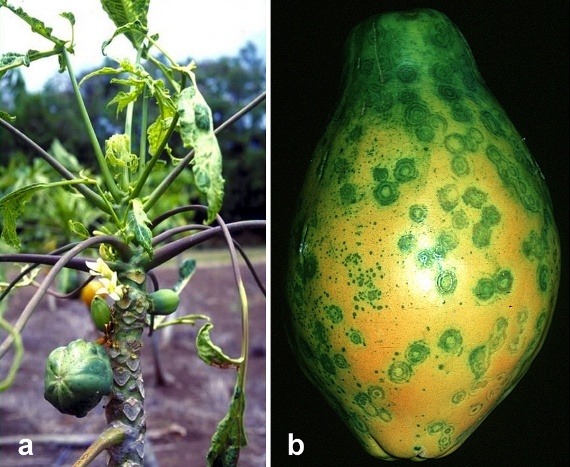
The commercial cultivation of papaya first began in the Hawaiian Islands. Around 1940, papaya plantations were first established on the island of Oahu, which provided a livelihood to many local farmers. In the 1950s, an outbreak of the papaya ringspot virus (PRSV) occurred in Oahu’s papaya plantations. This virus causes the stunting of trees, the deformation of leaves, the decline in their yields, and rounded spots in the infected fruit (see figure 7.5). Within a short period, the infected plant dies. Within a few years, the papaya plantations of Oahu were destroyed by the PRSV, so the industry moved to the big island of Hawaii, where the papaya industry flourished and expanded without any problems for the next four decades. In 1984, Hawaii’s papaya production reached its highest point (80.5 million pounds). However, around 1990, the PRSV appeared in Hawaii. As expected, the papaya plantations of Hawaii were devastated. By 1997, the papaya production declined up to 40 percent, Hawaii’s economy staggered, and many farmers’ livelihoods were destroyed.
Most of Hawaii’s papaya farmers were first-generation, less-educated immigrant Filipinos. Fortunately, Dennis Gonsalves, a Cornell University professor, grew up in Hawaii and knew the importance of papaya for the local economy. He noticed the PRSV when its outbreak began in Hawaii and started a research project to save the papaya plantations from PRSV. First, he tried to find a PRSV-resistant papaya variety so that trait could be transferred to the high-yielding productive papaya varieties using traditional plant breeding methods. But he didn’t succeed in finding such a variety. After this, Gonsalves identified less harmful strains of PRSV and experimented with them. He observed that if a papaya plant is first infected with a less aggressive strain, then the more harmful PRSV strain does not attack it. Until 1983, Gonsalves kept doing these experiments, but no permanent solution emerged for the disease’s practical management.
Around 1983, Roger Beachy made a kind of transgenic tobacco that was resistant to the tobacco mosaic virus (TMV). After entering a living cell, TMV (like most other viruses) makes replicas of its genome using the host cells’ resources, and then genome replication stops, and the synthesis of coat protein begins. The coat protein forms the virus’s outer shell; thus the viral genome gets packed one by one, and the progeny of TMV busts out by destroying the host cells. Beachy cloned the coat protein gene of TMV and then transferred it into tobacco plants. Beachy’s transgenic tobacco expressed high quantities of coat protein, so when TMV infected them, the coat protein present in the transgenic plant prevented replication of the viral genome, and infection was contained. Professor Gonsalves, inspired by Roger Beachy’s success, decided to introduce the PRSV’s coat protein gene in the papaya. In 1986, he successfully cloned this gene and transferred it within papaya plants with the help of a gene gun. As expected, the transgenic papaya showed resistance to the devastating PRSV. Field trials of these GE varieties, known as Rainbow and SunUp, began in 1992, and in late 1996, the US government approved transgenic papaya for commercial planting.
In 1998, Hawaiian farmers received Rainbow papaya seeds and the necessary advice free of cost, and Hawaii’s papaya plantations got a new life. Currently, a cooperative society sells seeds of PRSV-resistant papaya varieties in Hawaii to farmers at very low prices. For the past twenty-two years, the papaya of Hawaii has been selling globally and contributes $50 million annually to Hawaii’s economy. Gonsalves did not receive any grant from any government or private institution to make the GE papaya varieties. The only help he received was a small grant of $20,000 from Hawaii’s senator Daniel Inouye. He created the world’s first successful GE crop at a very low cost and made it available to farmers free of charge. In 2002, Gonsalves’s research team was awarded the Humboldt Research Award for their contribution to agriculture.
Later, others made virus-resistant GE squash, zucchini, potatoes, and plums (see table 7.1).
7.4.2 Roundup Ready Soybeans
Weed control has always been a challenge for farmers. Weeds are wild plants (sometimes closely related to crops), are well adapted to various environments, and are more resilient than crops. Weeds compete for nutrients, sunshine, and water with crops and negatively affect crop yields (20–40 percent). In traditional societies, farmers used to burn the weeds before sowing and then afterward spent time weeding. However, this is not an option for very large agricultural fields. After 1950, chemicals were discovered that can destroy the foliage. Initially, they were sprayed in the area before sowing to eradicate weeds. But these chemicals are hazardous for human health and have chronic and acute toxicity. Furthermore, many herbicides are carcinogens, and their use poses a risk of polluting water, air, and the food chain.
In 1970, John Franz, a scientist at the Monsanto Company, discovered herbicidal activity in glyphosate.[6]Furthermore, it was found that some soil bacteria can convert glyphosate into harmless substances within a few days. Hence it posed relatively less risk of environmental pollution compared to previously used chemicals. In 1974, it was marketed as an herbicide with the trademark Roundup. In the 1970s, Monsanto also invested in plant biotechnology and began developing herbicide-resistant transgenic crops. They already had information that soil microorganisms can metabolize glyphosate into harmless products. Soon scientists discovered a gene within a bacterium that coded for an enzyme capable of degrading glyphosate. Subsequently, this gene was introduced in soybean and corn to create Roundup Ready GE varieties that survive herbicide (glyphosate) sprays while all the weeds die. In 1996, a Roundup Ready soybean was approved by the FDA for its cultivation in the US. In this way, weed management became simpler, cheaper, and more manageable. Also, farmers can plant soybeans in a closer, tighter row and get higher yields. However, the second-generation soybeans are sterile, so farmers must buy new seeds every year from the seed company. This strategy was also used for creating herbicide-tolerant corn, rice, and many other crops. More than 50 percent of GE crops currently grown in the US and worldwide are herbicide resistant (see table 7.1).
Today, more than 90 percent of soybeans, corn, cotton, and mustard growing in the US are Roundup Ready varieties. Farmers around the world are increasingly relying on herbicides for weed management. Since 1996, when genetically engineered glyphosate-tolerant Roundup Ready crops were first released, glyphosate use has risen fifteenfold globally; these quantities are beyond the natural capacity of soil bacteria to metabolize glyphosate. Therefore, glyphosate and other herbicides have generated concern among the public and policymakers due to their harmful effects on human health and the environment.
7.4.3 Insect-Resistant Bt Crops
Most of us do not like to eat any fruit or vegetable that is infested by insects. We see the clean fruit and vegetables in the market because farmers have used plenty of pesticides. If pesticides are not sprayed, one-third of the yields of most crops would be lost due to pest infestation. However, pesticides have harmful effects on human health and pollute the environment.
Scientists have been looking for an alternative to pesticides for a long time. In 1901, a Japanese scientist, Shigetane Ishiwata, was investigating the cause of silkworms’ sudden deaths and found the bacterium Bacillus in dead larvae. He concluded that Bacillus is responsible for their deaths. In 1911, Ernst Berliner also noticed that the Mediterranean floor moths’ larvae were dying due to Bacillus’s presence. He named the bacterium Bacillus thuringiensis after the German city of Thuringia.
B. thuringiensis (or Bt) is a gram-positive soil bacterium that contains crystals of a delta-endotoxin protein that kills a broad category of insects. Upon ingestion, the crystal protein is cleaved into small fragments by proteases present in the alkaline environment of an insect’s gut. One of the fragments generated from the crystal protein acts as a toxin. This toxin kills insects by forming pores into the cell membranes of the insect midgut. From 1920 onward, European farmers began spraying the Bt bacteria to protect their crops from pests. In 1938, commercial spore-based formulations known as Sporine made it to the market in France. Even today, organic farmers spray a similar formulation of Bt bacteria on their crops.
In the 1980s, the Monsanto Company succeeded in cloning the “cry” gene from B. thuringiensis that codes for the crystal protein. Subsequently, this gene was optimized for high expression in plant cells to develop insect-resistant crops (e.g., cotton, maize, eggplant, and rice). When insects feed on Bt plants, the delta endotoxin reaches their intestine, and they die. Therefore, Bt crops can protect themselves from insects. In 1995, the US government approved Bt cotton (Bollgard cotton) and Bt corn for cultivation. Since then, these crops are being grown in the US and many other countries. These seeds of Bt varieties sold in the market are of the heterozygous F1 hybrid. Thus, farmers cannot save seeds of Bt crops for the following year’s sowing, because in the F2 generation, resistance and susceptibility traits segregate, and the insect-resistant traits get diluted in every subsequent generation. The farmers are required to buy these seeds every year, and their input costs increase.
The Bt crops do not eradicate insects completely, but their use could reduce the quantities of chemical pesticides manifold. The farm management is crucial for integrating these bioengineered crops and the chemical spray and keeping a sufficient refuge of non-GE crops to reduce the pests’ selection pressure. In the US, Brazil, Argentina, and so on, practicing integrated pest management (IPM) is easy because thousands of acres of the farm are run by a single person or company. But in other developing countries, farmers have smaller agricultural land holdings and do not have enough space for “refuge.” The farms are also surrounded by many neighboring farms.
Since living organisms constantly evolve, the protection offered by the Bt gene is not going to last for a long time, and eventually, it will be ineffective against insects. That is why scientists continue to make new versions of the Bt gene and search for other genes with similar properties. Thus it is expected that from time to time, scientists will release new GE varieties as the arms race between hosts and pests continues.
7.4.4 Golden Rice
The diet of poor people living in Asia, Africa, and Latin America mainly consists of rice and lacks fruits, vegetables, dairy, and meat. Consequently, millions of people have a huge deficiency of vitamin A in their bodies. According to a survey by the World Health Organization, more than 400 million people in twenty-six countries living on rice are deficient in vitamin A, due to which 500,000 children suffer from night blindness/blindness and 1 million children die annually.[7]
Vitamin A is essential for the proper development of humans and animals. It supports the healthy development of eyes, bones, and muscles and maintains adequate calcium levels and immunity. Humans cannot biosynthesize vitamin A of their own, but they get it from dairy, meat, and colorful vegetables and fruits. Red, orange, and yellow vegetables and fruits (e.g., sweet potatoes, carrots, oranges, and mangoes) contain provitamin A (β-carotene), which gets converted into vitamin A inside the human body.
For a long time, plant breeders have been searching for a rice variety in which β-carotene is found. In 1991, Ingo Potrykus, at the Institute of Plant Sciences of the Swiss Federal Institute of Technology, Zurich, started a project to bioengineer the β-carotene biosynthesis pathway in rice. In plants, the biosynthesis pathway consists of eight reactions, which are catalyzed by four enzymes. Rice contains most of the precursors for making β-carotene, but three out of four enzymes are nonfunctional. In 2000, Potrykus and his colleague Peter Beyer cloned two genes, phytoene synthase and lycopene cyclase, from the daffodil and the phytoene/carotene desaturase (crt1) gene from bacteria and then successfully introduced all three genes in rice plants to create the provitamin-rich variety by reconstructing the β-carotene biosynthesis pathway (see figure 7.6). One kilogram of this rice contains about eight milligrams of β-carotene, which gives them a golden hue, and thus this variety was named golden rice.[8] Once this basic strategy was successful, Potrykus signed a contract with the company Syngenta for enhancing the content of β-carotene and for large-scale experimental testing and handling. Syngenta produced an SGR2 variety of golden rice (~27mg β-carotene/kg) by replacing the phytoene desaturase in a daffodil with maize homolog. The Golden Rice Humanitarian Board has made golden rice free for farmers in developing countries with an annual income below $10,000. Later, with the help of research institutes of various countries, β-carotene biosynthesis pathways have also been enabled in many rice varieties (IR 64, Boro, PSB, RC82, etc.). After extensive testing, these GE rice varieties were found to be safe for human consumption. Unlike other GE crops, farmers who grow golden rice can save the seeds of the next year’s sowing. It took almost two decades before the golden rice was approved for farmers’ use. In 2018, the FDA, Health Canada, and Food Standards Australia New Zealand recognized the IRRI’s evaluation and approved the release of golden rice’s GR2E variety. Bangladesh and China have begun the cultivation of golden rice.

After rice, scientists started to work on biofortifying other crops using a similar strategy. For example, Maria Andrade, Robert Mwanga, and Jan Low created the golden sweet potato[9] with the support of global nonprofit agricultural research program HarvestPlus (https://www.harvestplus.org/), USAID, the Bill and Melinda Gates Foundation. Similarly, the golden cassava variety has also been developed, which has special importance for Africa.
Table 7.1 Herbicide-tolerant GE crops
| Crop | Herbicide | Year | Inventor |
|---|---|---|---|
| Canola | Glufosinate | 1995 | Bayer |
| Canola | glyphosate | 1999 | Monsanto |
| Cotton | Bromoxynil | 1994 | Calgene |
| Cotton | glyphosate | 1996 | Monsanto |
| Cotton | Sulphonylurea | 1996 | DuPont |
| Cotton | Glufosinate | 2003 | Bayer |
| Cotton | Dicamba | 2015 | Monsanto |
| Cotton | 2,4-Dichlorophenoxyacetic acid (2,4-D) | 2015 | Dow |
| Maize (corn) | Glufosinate | 1995 | AgrEvo GmbH |
| Maize | glyphosate | 1996 | Monsanto |
| Maize | 2,4-D | 2014 | Dow |
| Sweet corn | glyphosate | 2011 | Monsanto |
| Rice | Glufosinate | 1999 | AgrEvo GmbH |
| Soybean | glyphosate | 1994 | Monsanto |
| Soybean | Glufosinate | 1996 | Bayer |
| Soybean | Sulphonylurea | 2007 | DuPont |
| Soybean | Isoxaflutole | 2013 | Sygenta |
| Soybean | Mesotrione | 2014 | Sygenta |
| Soybean | Imidazolinone | 2014 | BASF |
| Soybean | 2,4-D | 2015 | Dow |
| Soybean | Dicamba | 2015 | Monsanto |
| Sugar beets | glyphosate | 2005 | Monsanto |
| Sugar beets | Glufosinate | 1998 | AgrEvo GmbH |
Table 7.2 Insect (pest)-resistant crops
| Crop | Transgene | Year | Inventor |
|---|---|---|---|
| Cotton | Bt | 1995 | Monsanto |
| Field corn | Bt | 1995 | Monsanto |
| Sweet corn | Bt | 1998 | Monsanto |
| Potato | Bt | 1995 | Monsanto |
| Soybean | Bt | 2016 | Monsanto |
Table 7.3 Virus-resistant crops
| Crop | Coat protein gene | Year | Inventor |
|---|---|---|---|
| Papaya (Rainbow and SunUp) | Ring Spot Virus | 1996 | Dannis Gonsalves (Cornell University, and ARS-USDA) |
| Plum | Plum pox virus | 2007 | USDA |
| Potato | Potato Virus Y | 1999 | Monsanto |
| Potato | potato leaf roll virus | 2000 | Monsanto |
| Squash (not widely grown) | zucchini yellow mosaic virus | 1994 | Asgrow |
| Squash (not widely grown) | watermelon mottle virus 2 | 1994 | Asgrow |
Table 7.4 Nutritional value improved crops
| Crop | Trait | Year | Inventor |
|---|---|---|---|
| Canola | High Iysine | 1994 | Calgene |
| Maize | High-Iysine | 2006 | Monsanto |
| Golden Rice | Rich in β-carotene (pro-vitamin A) | 2005 | Sygenta |
| Plenish Soybean | High oleic acid content (no trans fat) | 2010 | DuPont |
| Potato | Less acrylamide, when cooked/fried at a high temperature. | 2014 | J R Simplot Company |
| Arctic Apple | The GE Apples resist browning after being cut. | 2015 | Okanagan Specialty Fruits Inc. |
7.5 The Impact of GE Crops
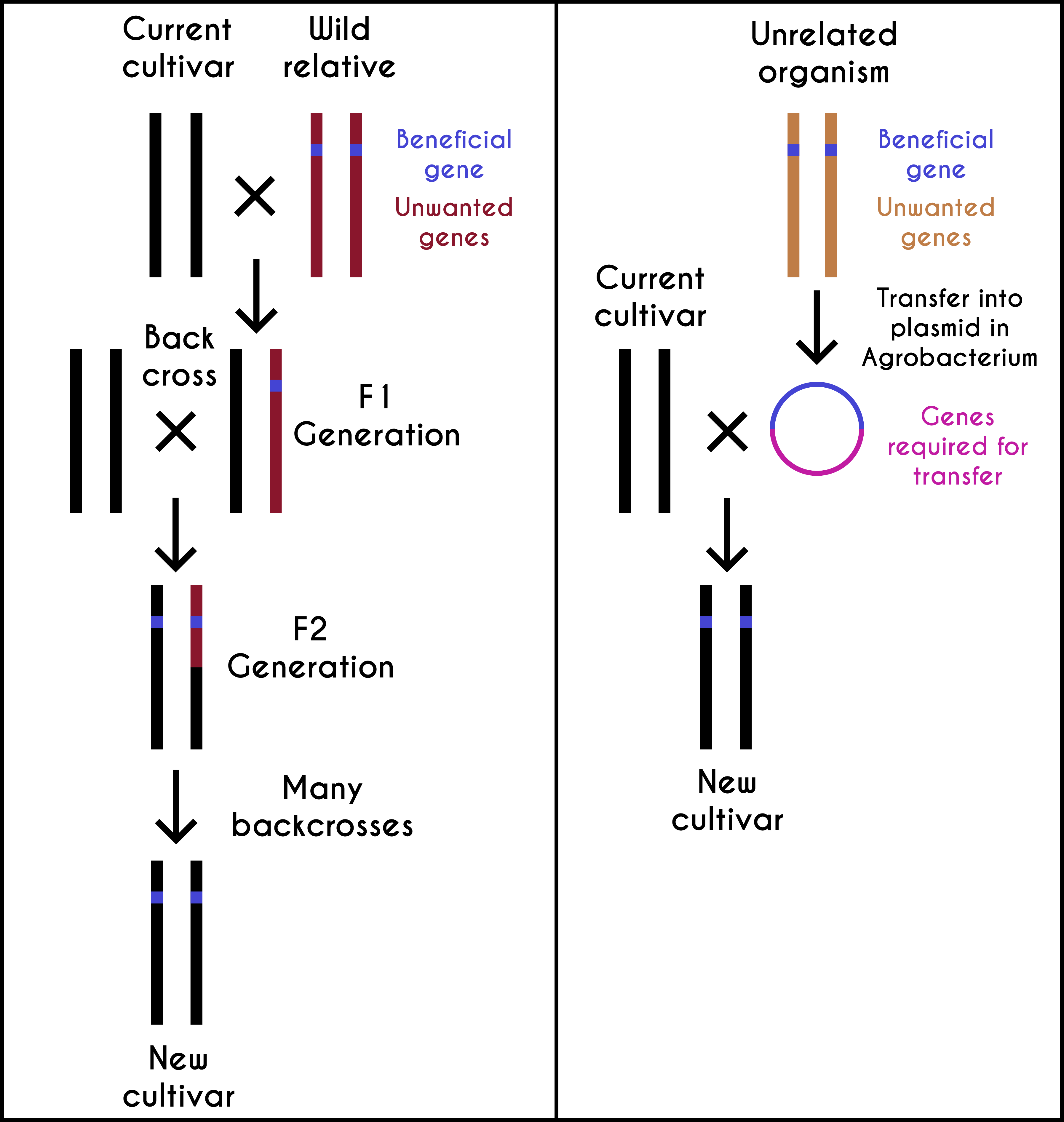
Humans have been selecting plants and animals for desired traits that suited their needs since the beginnings of agriculture 10,000 years ago. GE crops are the latest advancements made in this direction, with goals of achieving agricultural productivity and improving the available genetic stocks’ quality. After 1930, these objectives were achieved using traditional breeding. In the early twenty-first century, scientists genetically engineered crops by introducing useful genes from any living forms and/or by rationally designed genes. A GE crop may contain one or more transgenes from bacteria, viruses, animals, or plants. However, more than 99 percent of it is similar to its mother plant (see figure 7.7). In general, crop plant species contain 30,000–50,000 genes, and less than five genes are introduced in any given GE crop. In this sense, GE crops are closer to the domesticated varieties.
Interestingly, people have adopted more than 1,000 high-yielding varieties (generated by traditional breeding), but GE crops have aroused significant anxiety around the world and have faced consistent opposition globally. There has been a constant debate about the technology and impacts of GE crops. The first and foremost concern is if GMO food is safe. GE crops undergo extensive safety trials before they are released for cultivation and are approved by the FDA. The data from various GE crops’ safety trials in the last twenty-five years show that GMO food is safe. In 2016, the US National Academy of Sciences published a review of GE crops prepared by an independent committee of eighteen experts. This 600-page report (1) concluded that there is no evidence that anyone has had a problem digesting GMO foods so far. However, people have raised issues related to allergies and the presence of toxins in GMO food. Overall, GMO food is deemed safe by the experts. Today most experts, scientific institutions, and governments agree that GMO food does harm human health. According to an estimate, millions of people in twenty-eight countries have been consuming GMO food products daily for the past twenty-five years, and so far, none died due to eating such foods (1).
Contrary to what the experts have revealed, there is increasing anxiety among consumers about GMO products. This issue has not been settled for the people yet. Many nongovernmental organizations, consumer groups, and ordinary people worldwide are worried about the safety of consuming GMO food. No one has currently investigated GMO products’ long-term effects on humans; therefore, both opponents and supporters of GMOs can’t say anything definitively on the matter. For a long time, consumer groups had been advocating to clearly label products that are GMO, since in the US, this had not been required by law. Under consumers’ pressure, the labeling of GMO (and more of non-GMO) products appeared increasingly, and in 2016, US Congress passed a law requiring the labeling of GE food (the bioengineered). Hence, the data on the long-term effects of GMO food on human and animal health may become available in the future.
Another issue is how much more productive GE crops are than the high-yielding breeds developed during the green revolution. In the last twenty-five years, the claims made about GE crops’ increased productivity have not been fully met. The 2016 National Science Academy report (1) stated that the productivity of GE crops grown from 1990 to 2015 is not significantly higher than the high-yielding varieties of the green revolution era. Overall, the GE crops made the farmers less prosperous than the green revolution because agriculture’s input cost (for buying seeds, fertilizer, irrigation, pesticides, fungicides, herbicides, etc.) is very high and leaves a small margin for profit. Compared to the green revolution era, today’s farmers have smaller and less fertile farms in developing countries, and government assistance and subsidies have declined.
Did GMO technology improve the quality of food? Without a doubt, this technology has created many nutrient-rich crops—like golden rice, cassava, and sweet potato; high oleic acid–containing canola; Plenish soybeans; and so on—and offers an opportunity for further improvements. There is tremendous scope for solving malnutrition by using GMO technology. In addition, the genetic engineering of model and crop plants has provided a conceptual understanding of gene functions and many fundamental biological processes. Besides genetics and genomics, immense progress has been made in vitro tissue culture technology. Together, the new knowledge and advancement in tools served as a foundation for next-generation CRISPR/Cas9 gene-editing technology that has a high potential for developing new biofortified and stress-tolerant varieties of crops.
At the beginning of the twenty-first century, GE crops were presented as a promising alternative to a non–environmentally friendly industrial farming system. But looking at the last twenty-five years, it is clear that GE crops have become an integral part of it. The folks who oppose GMOs argue that most GE crops ensure profit to corporations by encouraging increased use of pesticides, weedicides, fungicides, and fertilizers. To date, more than 50 percent of GE crops are herbicide resistant. The use of glyphosate and other herbicides has exceeded the safety threshold and poses threats to human health and the environment. The continuous spray of herbicides has led to the rise of superweeds that are increasingly adapting to various herbicides and require more potent mixtures (including two or more chemicals) or higher doses for effective control. Thus GM crop-based weed management is not sustainable. Additionally, the reliance on both high-yielding and GM crops on heavy inputs of energy and fertilizers also needs careful reevaluation.
7.6 Toward a Sustainable Future
The FAO projects the global population to grow to 9.7 billion by 2050. The biggest challenge of the twenty-first century is to keep the productivity of crops sustainable without causing further destruction to the environment and depletion of natural resources. Despite its overall efficiency, the current industrial agricultural system relies heavily on natural resources and energy. For example, half of the world’s habitable land and more than 70 percent of global freshwater withdrawals are used for agriculture. Also, agriculture is a significant source of global pollution: 26 percent of global greenhouse gas emissions[10] and 78 percent of the global ocean and freshwater eutrophication is caused by agriculture.[11] Agriculture is the prime source of greenhouse gases, including carbon dioxide (CO2) and nitrous oxide (N2O), which cause the depletion of the stratospheric ozone layer and global warming. Therefore, the agricultural production systems have a big impact on global climate change, natural resources, and all living beings’ health.
It is important to note that a continuous increase in agriculture productivity alone cannot solve hunger in the world. Natural resources are finite, and their overexploitation will have disastrous consequences. Therefore, we must explore other avenues for bringing efficiency to ensure food security. Today, enough grain is being produced worldwide to meet the need of 10 million people. Theoretically, if food is distributed equally among all the people of the world, then 75 percent of it is enough to feed the entire population of the world, and one-fourth can be saved, although achieving such equity at the global level is difficult to achieve due to various sociopolitical reasons. Nonetheless, it suggests that there are additional avenues where progress can be made beyond agriculture production.
It is noteworthy that more than 40 percent of all food produced worldwide is wasted in processing, transportation, supermarkets, and kitchens. Many fruits and vegetables are discarded because of their lack of aesthetic appeal or uniformity. Moreover, readymade, cheap, and readily available food requires extra production to compensate for its regular trashing as it expires on the supermarkets’ shelves. When food is wasted, all the resources to grow, process, package, and transport are wasted along with it. Besides, the waste is a source of significant greenhouse gases. Overall, the current agricultural production systems and consumers’ lifestyles put a burden on natural resources, the environment, and energy. If consumers do not change their habits, then the agricultural production system will remain in its current form. Both farmers and consumers will have to strive together to build a new sustainable agricultural production system.
In past decades, the gradual progress in plant genetics, genomics and breeding, precision agriculture, organic agriculture methods, and growing consumer awareness has been paving the way for the agricultural production system’s future. Better management of agriculture, natural resources, the environment, and human resources is required in the agriculture sector. The formula for successful and sustainable agriculture will have elements from industrial farming, GM technology, precision agriculture, and organic farming. Equally important is to foster a culture of conscious consumption and change in consumers’ behavior to create new, sustainable, just, diverse, and healthy agricultural production systems. The issue of agriculture has never been limited to production. It has given rise to human civilization and its polytheistic cultures. Change in agriculture is intricately associated with a change in human society. Only a better society can build a better agricultural system.
References
National Academies of Sciences, Engineering, and Medicine. (2016). Genetically engineered crops: Experiences and prospects. National Academies Press. https://doi.org/10.17226/23395 (↵ Return 1) (↵ Return 2) (↵ Return 3)
Further Readings
Benbrook, C. M. (2016). Trends in glyphosate herbicide use in the United States and globally. Environ. Sci. Eur., 28, 3.
Beyer, et al. (2000). Engineering the provitamin A (-carotene) biosynthetic pathway into (carotenoid-free) rice endosperm. Science, 287(5451), 303–5. https://doi.org/10.1126/science.287.5451.303
Gillam, C. (2018, May 8). Weedkiller products more toxic than their active ingredient, tests show. The Guardian. https://www.theguardian.com/us-news/2018/may/08/weedkiller-tests-monsanto-health-dangers-active-ingredient
Gonsalves, D. (1998). Control of papaya ringspot virus in papaya: A case study. Annu. Rev. Phytopathol, 36, 415–37.
Jenkins, M. (2018). Food fight: GMOs and the future of the American diet. Avery.
Oerke, E. C. (2006). Crop losses to pests. J. Agric. Sci., 144, 31–43. https://doi.org/10.1017/S0021859605005708
Pingali, P. L. (2012). Green revolution: Impacts, limits, and the path ahead. PNAS, 109, 12302–8. https://doi.org/10.1073/pnas.0912953109
Pollan, M. (2006). The omnivore’s dilemma: A natural history of four meals. Penguin.
Poore, J., & Nemecek, T. (2018). Reducing food’s environmental impacts through producers and consumers. Science, 360(6392), 987–92.
Searchinger, T., et al. (2018). Creating a sustainable food future—a menu of solutions to feed nearly 10 billion people by 2050. World Resources Institute. https://www.wri.org
Tian, H., et al. (2020). Comprehensive quantification of global nitrous oxide sources and sinks. Nature, 586(7828), 248–56. https://doi.org/10.1038/s41586-020-2780-0
Watson, J. D. (1968). The double helix: A personal account of the discovery of the structure of DNA. Atheneum.
- An organism containing a transgene is known as a genetically modified organism (GMO). ↵
- Not all GE crops are in cultivation today. Only a few are grown in the US. To see the updated information on GE crops, visit the FDA’s website: https://www.fda.gov/food/consumers/agricultural-biotechnology. ↵
- The animals have additional DNA inside mitochondria; plants have DNA inside chloroplast and mitochondria. These are identified separately as organellar genomes. ↵
- Have a look at the Tree of Life at http://tolweb.org/tree, and see the Tree of Life segment of BBC One’s “Charles Darwin and the Tree of Life,” narrated by David Attenborough, at https://youtu.be/H6IrUUDboZo. ↵
- See https://www.fda.gov/food/consumers/agricultural-biotechnology. ↵
- Glyphosate (N-[phosphonomethyl] glycine) was discovered by Henri Martin in 1950. ↵
- For current data, visit https://ourworldindata.org/micronutrient-deficiency. ↵
- The story of golden rice, as told by Ingo Potrykus, is available at http://www.agbioworld.org/biotech-info/topics/goldenrice/tale.html. ↵
- Learn more about the golden sweet potato at https://www.worldfoodprize.org/en/laureates/20102019_laureates/andrade_mwanga_low_and_bouisold_template/?nodeID=88375&AudienceID=1&preview=1. ↵
- Agriculture is the prime source of greenhouse gases, including carbon dioxide (CO2), nitrous oxide (N2O), and methane, which cause the depletion of the stratospheric ozone layer and global warming. N2O has 300 times the warming potential of CO2. Usually, N2O is destroyed in the upper atmosphere by solar radiation. But current human activities (e.g., agriculture and livestock) have significantly increased N2O emissions and caused the perturbation in its natural cycle, resulting in its accumulation in the atmosphere. A comprehensive study by the Global Carbon Project and the International Nitrogen Initiative suggests that agriculture has contributed to a 70 percent rise in the global N2O levels. See the Global Carbon Project website for more details: https://www.globalcarbonproject.org/nitrousoxidebudget. ↵
- See https://ourworldindata.org/environmental-impacts-of-food. ↵

